Namespace pipelines¶
This section covers the following:
A brief introduction to namespaces and modular pipelines
How to convert the existing spaceflights project into a namespaced one
Adding namespaces to modular pipelines unlocks some sophisticated functionality in Kedro
You are able to instantiate the same pipeline structure multiple times, but provide different inputs/outputs.
You can unlock the full power of micro-packaging.
You can de-clutter your mental model with Kedro-Viz rendering collapsible components.
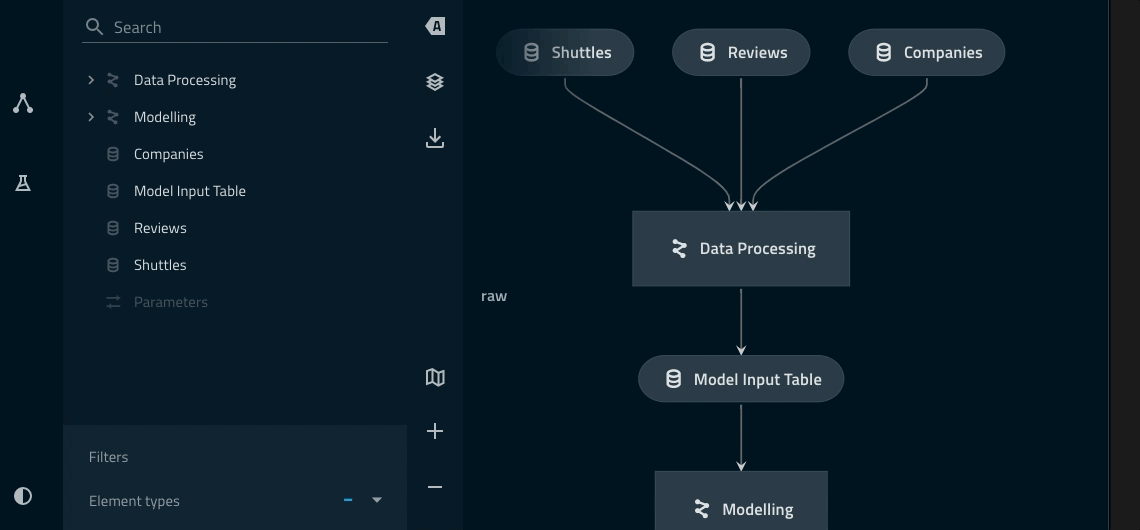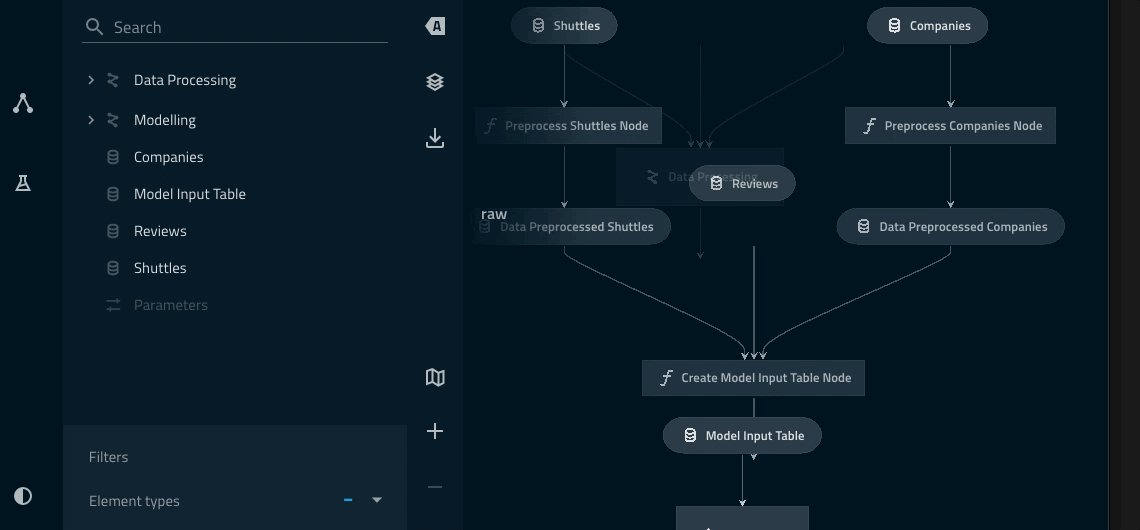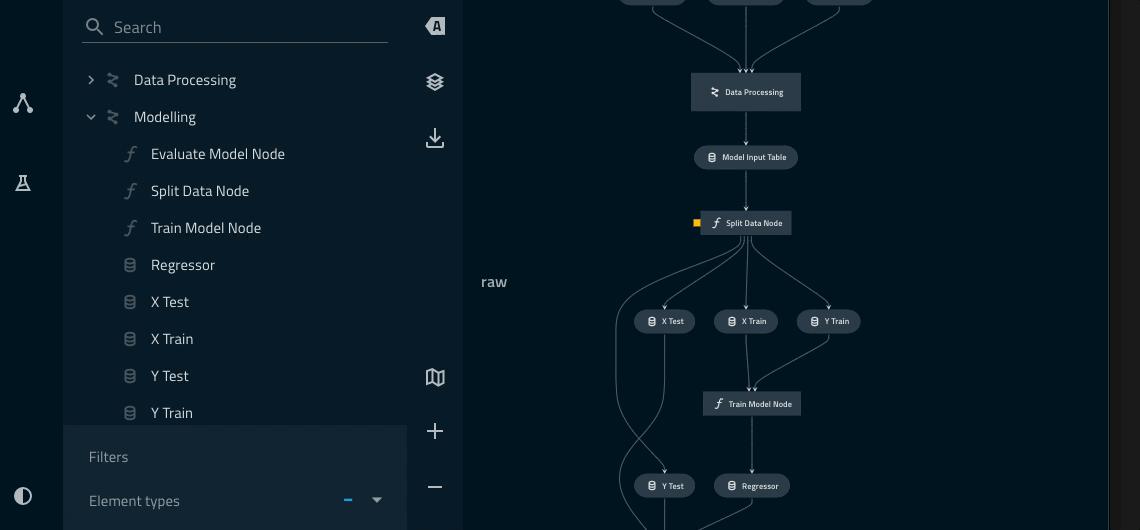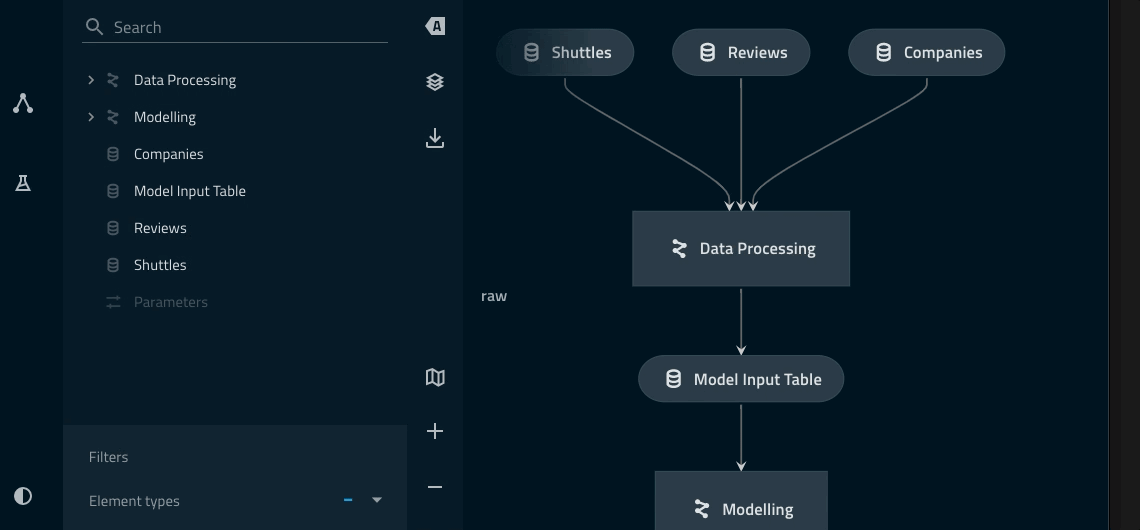
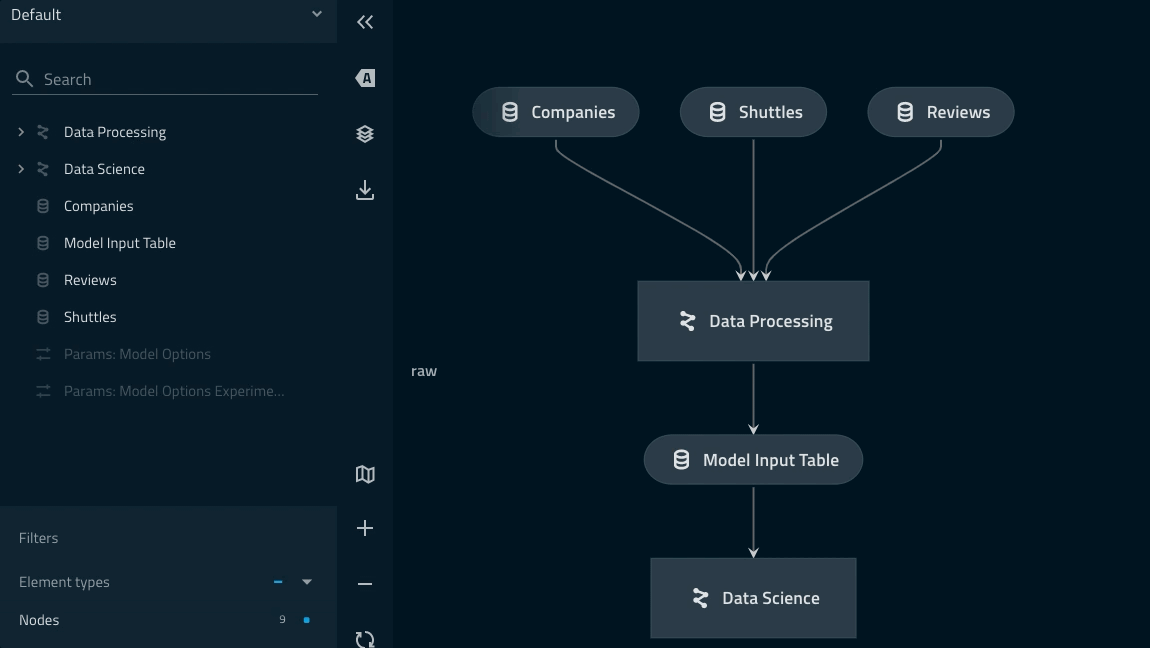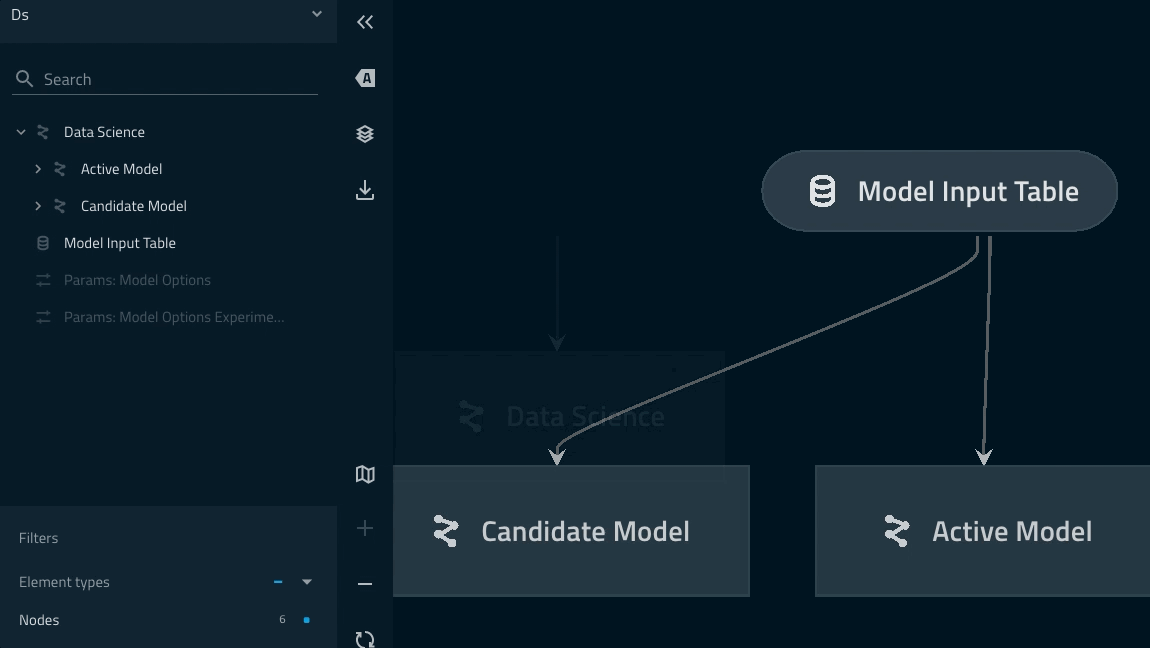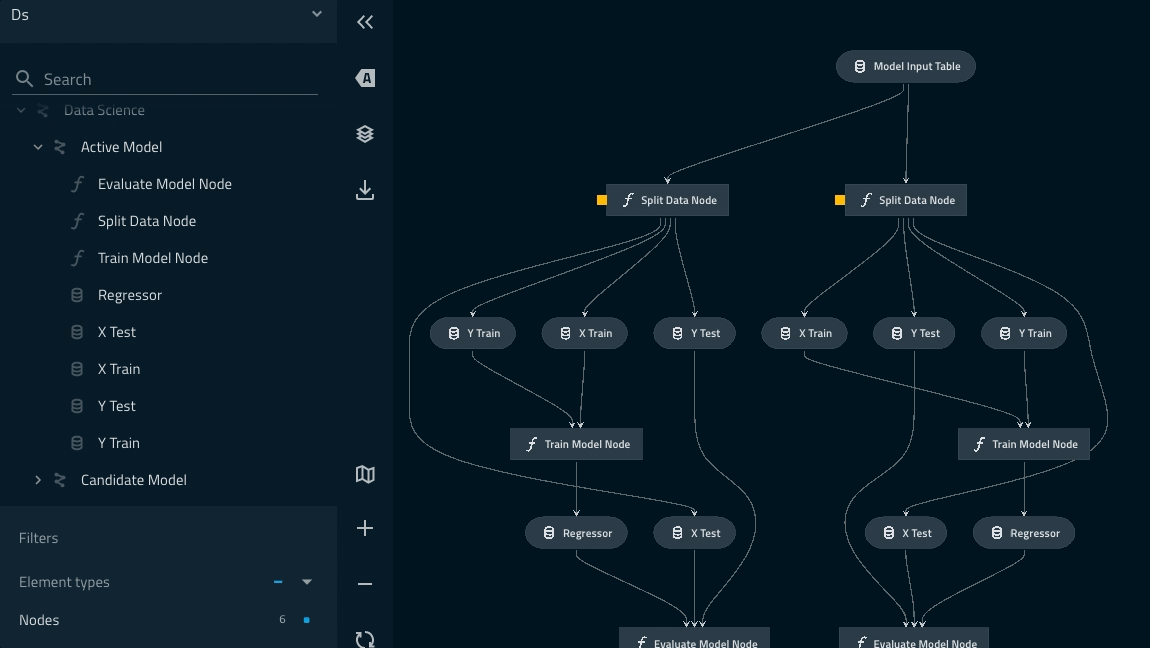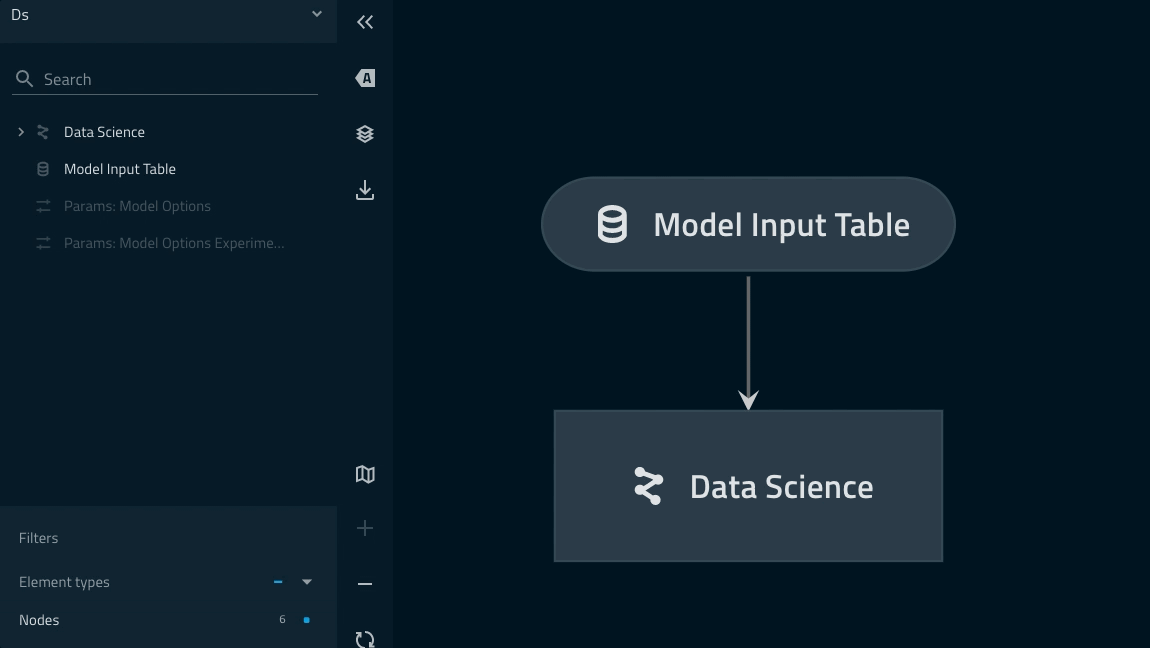
Adding a namespace to the data_processing pipeline¶
Update the code in
src/kedro_tutorial/pipelines/data_processing/pipeline.pyusing the snippet below.This introduces the lowercase
pipeline()method imported fromkedro.pipeline.modular_pipelineClick to expand
from kedro.pipeline import Pipeline, node from kedro.pipeline.modular_pipeline import pipeline from kedro_tutorial.pipelines.data_processing.nodes import ( preprocess_companies, preprocess_shuttles, create_model_input_table, ) def create_pipeline(**kwargs) -> Pipeline: return pipeline( [ node( func=preprocess_companies, inputs="companies", outputs="preprocessed_companies", name="preprocess_companies_node", ), node( func=preprocess_shuttles, inputs="shuttles", outputs="preprocessed_shuttles", name="preprocess_shuttles_node", ), node( func=create_model_input_table, inputs=["preprocessed_shuttles", "preprocessed_companies", "reviews"], outputs="model_input_table", name="create_model_input_table_node", ), ], namespace="data_processing", inputs=["companies", "shuttles", "reviews"], outputs="model_input_table", )
Why do we need to provide explicit inputs and outputs?¶
When introducing a namespace you must tell Kedro which inputs/outputs live at the ‘edges’ of the namespace
Failing to do so in this situation causes Kedro to think that
companiesanddata_processing.companiesare two different datasets.Highlighting ‘focus mode’ Kedro-Viz highlights how the explicitly declared inputs/outputs live outside of the collapsed part of the namespace.
model_input_tablein particular is shared across both pipelines and thus needs to be outside of theData Processingnamespace.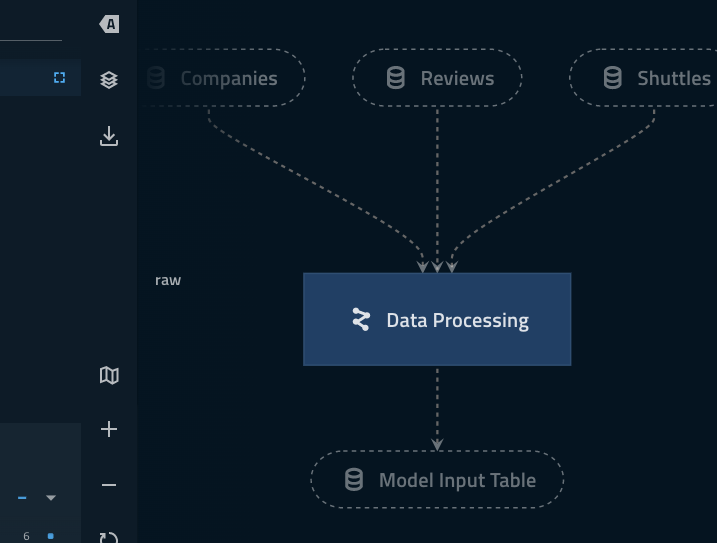
As an experiment, remove the explicit inputs and outputs from your
namespaced_pipelineand see how this creates a disjointed pipeline visualisation
Adding namespaces to the data_science pipeline¶
In this section we want to add some namespaces in the modelling component of the pipeline and also highlight the power of instantiating the same modular pipeline multiple times with different parameters.
Add some more parameters to the bottom of
conf/base/parameters/data_science.ymlusing this snippet:model_options_experimental: test_size: 0.2 random_state: 8 features: - engines - passenger_capacity - crew - review_scores_rating
Update you catalog to reflect the new namespaced outputs. Replace the
regressorkey with the following two new dataset keys inconf/base/catalog.yml:active_modelling_pipeline.regressor: type: pickle.PickleDataSet filepath: data/06_models/regressor_active.pickle versioned: true layer: models candidate_modelling_pipeline.regressor: type: pickle.PickleDataSet filepath: data/06_models/regressor_candidate.pickle versioned: true layer: models
Update the code in
pipelines/data_science/pipeline.pyusing the snippet below.Click to expand
from kedro.pipeline import Pipeline, node from kedro.pipeline.modular_pipeline import pipeline from .nodes import evaluate_model, split_data, train_model def create_pipeline(**kwargs) -> Pipeline: pipeline_instance = pipeline( [ node( func=split_data, inputs=["model_input_table", "params:model_options"], outputs=["X_train", "X_test", "y_train", "y_test"], name="split_data_node", ), node( func=train_model, inputs=["X_train", "y_train"], outputs="regressor", name="train_model_node", ), node( func=evaluate_model, inputs=["regressor", "X_test", "y_test"], outputs=None, name="evaluate_model_node", ), ] ) ds_pipeline_1 = pipeline( pipe=pipeline_instance, inputs="model_input_table", namespace="active_modelling_pipeline", ) ds_pipeline_2 = pipeline( pipe=pipeline_instance, inputs="model_input_table", namespace="candidate_modelling_pipeline", parameters={"params:model_options": "params:model_options_experimental"}, ) return ds_pipeline_1 + ds_pipeline_2
Let’s explain what’s going on here¶
Modular pipelines allow you instantiate multiple instances of pipelines with static structure, but dynamic inputs/outputs/parameters.
pipeline_instance = pipeline(...)
ds_pipeline_1 = pipeline(
pipe=pipeline_instance,
inputs="model_input_table",
namespace="active_modelling_pipeline",
)
ds_pipeline_2 = pipeline(
pipe=pipeline_instance,
inputs="model_input_table",
namespace="candidate_modelling_pipeline",
parameters={"params:model_options": "params:model_options_experimental"},
)
The
pipeline_instancevariable is our ‘template’ pipeline,ds_pipeline_1andds_pipeline_2are our parametrised instantiations.
The table below describes the purpose of each keyword arguments in detail:
Keyword argument |
|
|
|---|---|---|
|
Declaring |
Same as ds_pipeline_1 |
|
No outputs are at the boundary of this pipeline so nothing to list here |
Same as ds_pipeline_1 |
|
Inherits defaults from template |
Overrides provided |
|
A unique namespace |
A different unique namespace |
Nesting modular pipelines¶
Modular pipelines can be nested an arbitrary number of times
This can be an effective pattern for simplifying you mental model and to reduce visual noise
Namespaces will be chained using the
.syntax just youimportmodules in PythonYou can quickly wrap your two modelling pipeline instances under one ‘Data science’ namespace by adding the following to your
pipelines/data_science/pipeline.pyreturn statement:... return pipeline( pipe=ds_pipeline_1 + ds_pipeline_2, inputs="model_input_table", namespace="data_science", )
This renders as follows: